Request a Quote
NJ Bio’s biologists are experts in the design and optimization of protein production using both bacterial and mammalian cells. Our team will identify potential risks and engineer personalized expression systems to get the job done. To get started, all that is needed from our clients is the amino acid sequence of the desired protein.
What we do with your amino acid sequence:
- Optimize the sequence
- Synthesize the gene
- Clone the gene into a vector
- Produce large-scale plasmid DNA
- Optimize protein expression
Protein Production:
- Bacterial Protein Production
- Tagged or untagged versions
- Variety of vectors
- Solubility enhancing fusions
- Purification method development
- Mammalian Protein Production
- Protein production in HEK293 or CHO cells
- Protein production using stable transient pools
- Purification using FPLC
QC/Analysis:
- SDS Page
- Western Blot
- HPLC
- Analytical SEC
- Mass Spectrometry
- Polishing SEC
- Endotoxin Level Measurement
Additional Services:
- 15N Isotope labeling for protein-NMR
Case Studies for Protein Production
1. Production of THIOMAB in Mammalian Cells
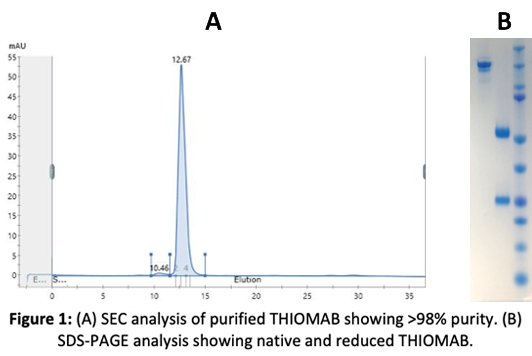
The received THIOMAB DNA sequence was optimized for codon usage, the gene synthesized and cloned it into a vector under a strong CMV promoter and a strong secretory sequence. The cells were transiently transfected and several clones were tested for protein expression. Protein expression was optimized using different conditions, such as temperature and different H:L chain ratios.
The produced THIOMAB protein was purified by Protein A affinity chromatography. Our transient transfection yielded the maximum amount of protein, which was further analyzed for purity and aggregation by SEC.
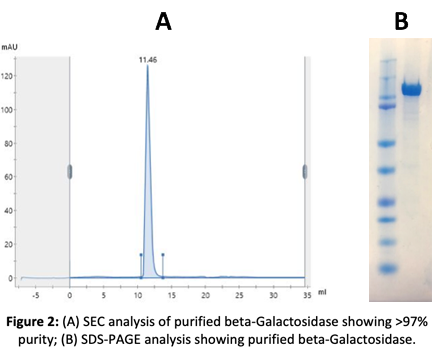
2. Production of Beta-Galactosidase in E. coli Bacteria
Our client provided the amino acid sequence of cold-active beta-Galactosidase. The sequence was optimized for codon usage in E. coli. The gene was then synthesized and cloned into a bacterial vector and protein production was optimized to maximize expression and solubility.
The protein was purified using Ni+ NTA protocols and was then subjected to functional and activity assays. After a final polishing step by SEC, the exact mass of beta-galactosidase was confirmed by mass spectrometry (TOF).